Get up to date with the latest HitGen articles and join us in the events
DNA-encoded library (DEL) technology represents a revolutionary method in drug discovery. Unlike conventional approaches, DEL enables unparalleled exploration of chemical space, leading to the rapid identification of novel potent compounds. Although the importance of DEL screening in hit discovery has been well recognized, the power of DEL technology in compound optimization has only been appreciated recently. Researchers can systematically explore chemical space around existing compounds using DEL technology, generating diverse analogs and derivatives (Figure 1). This approach is particularly powerful in identifying analogs with enhanced properties, such as increased potency, selectivity, or reduced toxicity. The ability to utilize DELs for compound optimization highlights their versatility and impact in drug discovery endeavors.
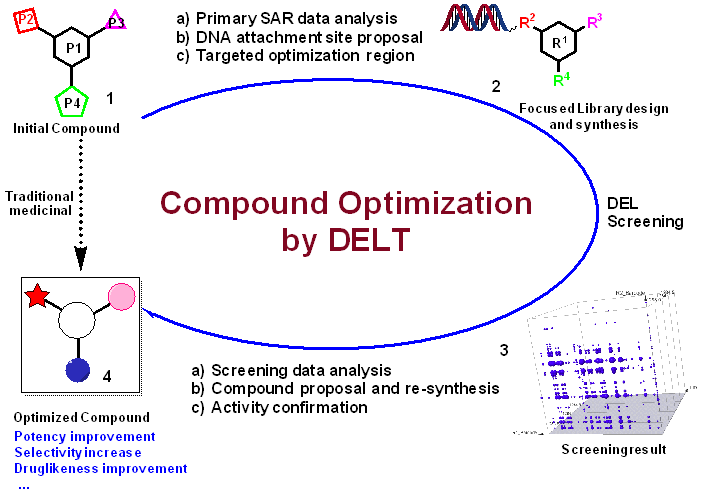
Figure 1: DELs for compound optimization
Fragment expansion using focused DELs
Fragment-based drug discovery (FBDD) is one of the most well-developed approaches for drug discovery starting from small, low-affinity compounds. These low-affinity fragments pose a challenge in evolving them into compounds with desirable affinities. The most widely accepted approach for enhancing the potency of fragments involves iterative fragment growing, a process that can be intricate and time-consuming.
In 2022, a collaboration between HitGen and Vernalis teams has led to the development of PAC-FragmentDEL, a novel approach involving the photoactivated covalent capture of DNA-encoded fragments (RSC Med. Chem., 2022, 13, 1341-1349). This method facilitates the rapid identification of fragments binding to specific targets. This approach only requires a small amount of tagged protein (250 pmol per sample), therefore reducing the quantity of biological targets necessary for screening. Moreover, the approach is less time-consuming, allowing the identification of hits within a few weeks including library selection, sequencing, and comprehensive analysis. Importantly, PAC-FragmentDEL can incorporate fragments with lower solubility, expanding the range of compounds compared to traditional methods. Additionally, the fragments identified through PAC-FragmentDEL have clear DNA-tagging points, providing valuable insights into the interactions between the fragments and their respective target proteins.
Recently, an innovative strategy for fragment hit optimization utilizing poised DNA-encoded chemical libraries was proposed (Chem. Sci., 2023, 14, 8288-8294). These poised libraries allow researchers to systemically investigate chemical space, facilitating the swift transformation of fragments into potent hit/lead compounds. In contrast to this method, the fragments identified through PAC-FragmentDEL possess established DNA-tagging points and functional groups, serving as reaction headers for the construction of fragment-focused DNA-encoded libraries (Figure 2).
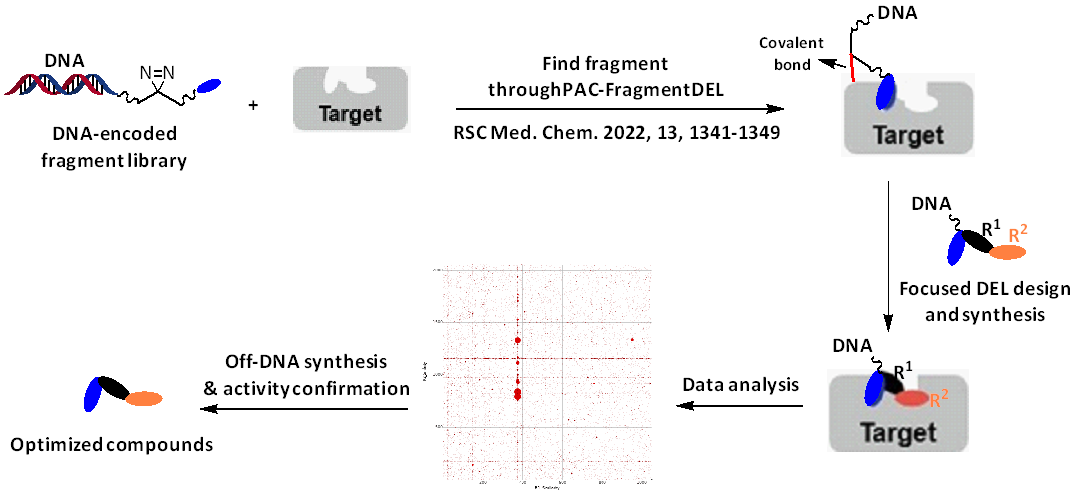
Figure 2: PAC-FragmentDEL selections and fragment expansion using focused DELs
Aiming to generate potent protein kinase PAK4 inhibitors, we incorporated diverse in-house DEL intermediates into the fragments of PAK4 identified through PAC-FragmentDEL (RSC Med. Chem., 2022, 13, 1341-1349), resulting in fragment-focused DELs containing over 100 million encoded compounds within two weeks. These resultant libraries then underwent screening against PAK4 utilizing traditional affinity-based selection. Following comprehensive data analysis, off-DNA synthesis, and activity confirmation, multiple series of hit compounds exhibiting low hundreds of nM activity were identified. The entire process spanning from library design and synthesis, library selection and data analysis, as well as off-DNA compound synthesis and subsequent validation only took 4-5 weeks (Figure 3).
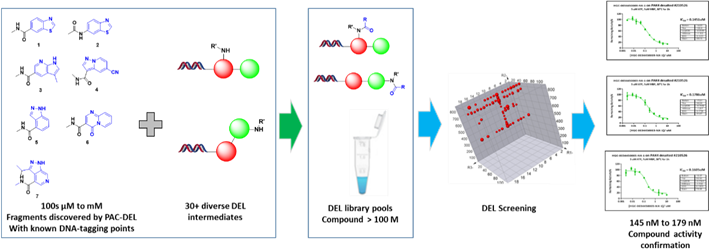
Figure 3: fragment expansion using focused DELs
Optimization of known hit/lead compounds using focused DELs
DEL technology enables systematic exploration of the chemical space surrounding known hit/lead compounds, facilitating the generation and screening of extensive and diverse focused compound libraries. This approach aims to enhance the potency, selectivity, or druglikeness of these compounds.
At HitGen, DEL technology has been employed in various projects for compound optimization. For instance, to enhance the selectivity of a reported compound, we designed two focused DNA-encoded libraries comprising millions of compounds, which involved confirming DNA-tagging points, designing and synthesizing core scaffolds, and controlling physicochemical properties (Figure 4). This whole process could be facilitated by automated instrumentation that allowed multiple DEL selection experiments to run simultaneously. Furthermore, the "Smart Selection Campaign" strategy was utilized, prioritizing DEL screening selectivity for proteins (wild type, mutations, etc.) from the starting point. By filtering out overlapping binders, specific inhibitor series were identified and subsequently validated, demonstrating exceptional selectivity.
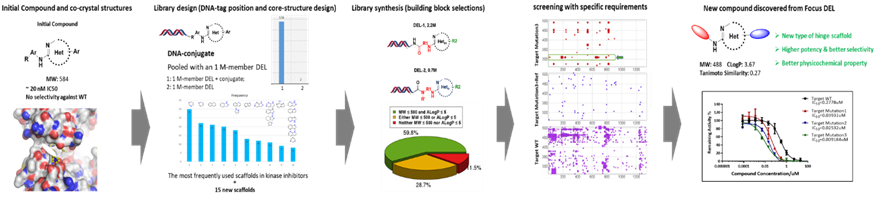
Figure 4: hit/lead compound optimization using focused DELs
DEL for compound optimization at HitGen
DEL technology exhibit significant potential applications beyond the well-established novel hit discovery, proving high values for compound optimization. Unlike traditional methods, this approach avoids the iterative cycles of Design-Make-Test-Analyze (DMTA), streamlining the optimization process significantly. Through the design and synthesis of focused DELs, HitGen achieves the following objectives:
Ø Supporting fragment-to-lead optimization: HitGen combines PAC-FragmentDEL with subsequent focused fragment-expansion DEL techniques to facilitate the optimization process from fragments to lead compounds.
Ø Optimizing known hit or lead compounds: HitGen employs focused DELs to optimize existing hit or lead compounds, enhancing their potency, selectivity or druglikeness.
We use cookies to provide a better web experience.
By using our site, you acknowledge our use of cookies and please read our Cookie Notice for
More information