One-stop Technology Platform to Accelerate Drug Discovery and Development
Affinity selection-mass spectrometry (ASMS) is an efficient and convenient high throughput screening approach for separating and identifying potential ligands from a complex mixture. It assesses the binding of candidate molecules to immobilized protein, whereas a mixture is incubated with target protein and then the potential ligands bonded to the receptor will be eluted and detected by MS after removal of the supernatant containing the components without binding. The advantage of this technique is all the possible binders are detected as a whole without separation. It accelerates the screening process and reduces the cost for bio affinity screening. ASMS is a valuable complement to traditional drug discovery technologies, especially in the discovery for hit identification.
The encoded tags of DEL after affinity screening is presented as signal enrichment cubic view for feature selection. The chemical structures of selected features from the cube are deducted according to their hit-index, and proposed for resynthesis for hit identification. Instead of hit proposal with deducted structures, ASMS with on-DNA validation is a powerful alteration. The on-DNA conjugates of deducted chemical structures are resynthesized with the same condition as in DEL production. This mixture of resynthesized products and their byproducts are incubated with the same immobilized protein using affinity selection assay for selection of less components that are binding to target protein. These components are then identified using MS, and assigned to species of the on-DNA conjugate.
ASMS is used in on-DNA validation in order to select possible binders with high confirmation rate. This approach takes both products and byproducts into consideration by mimicking the reactions during DEL production. We are also exploring different kinds of cleavable linkers, which will maintain the same conditions in DEL production but also avoid affections from their DNA tags during affinity selection.
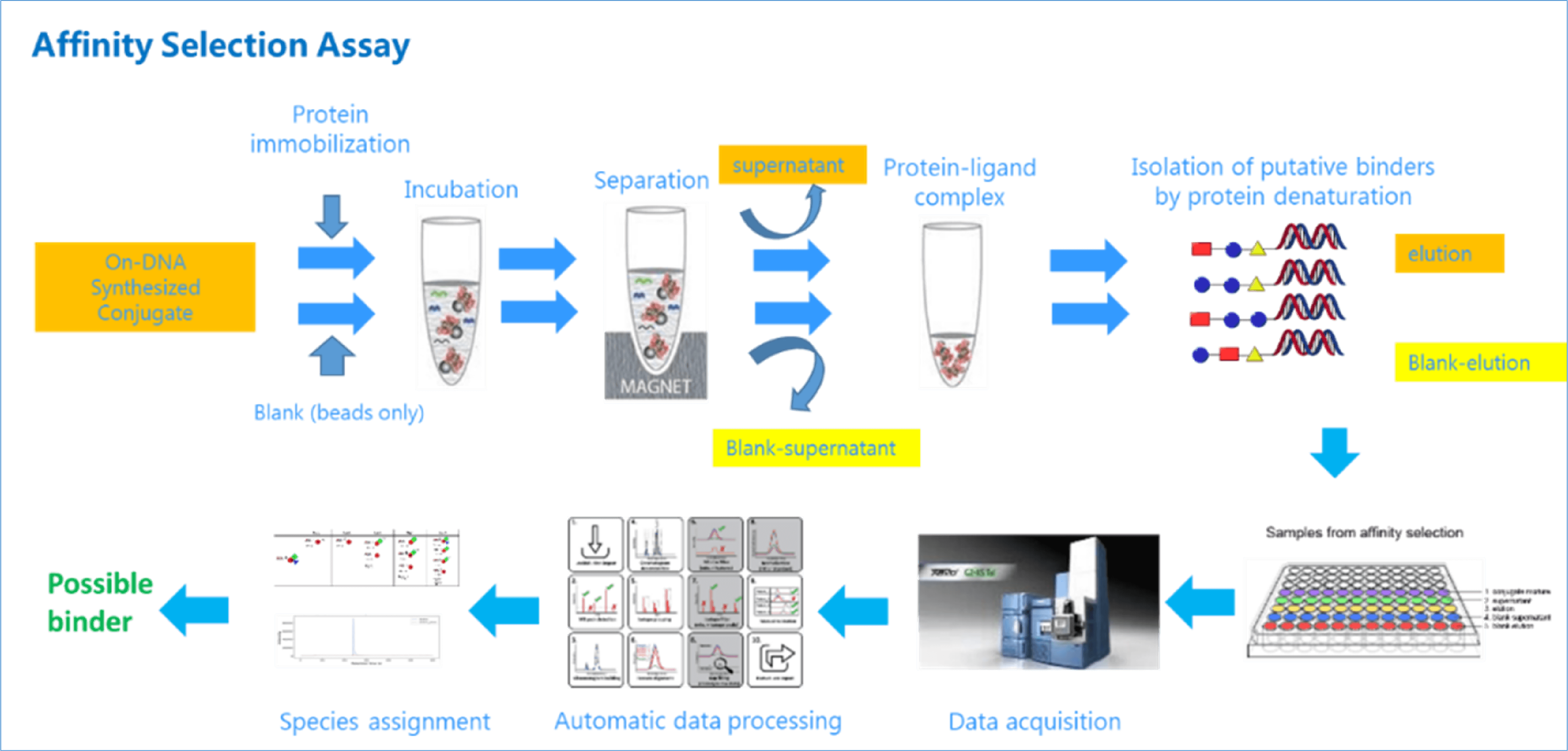
Workflow of ASMS
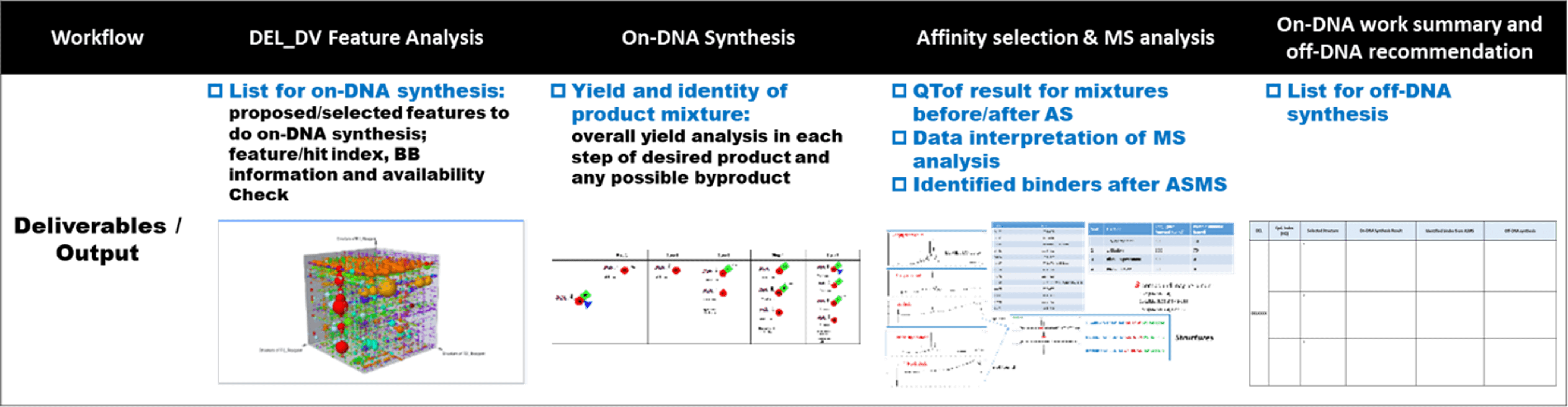
Our on-DNA ASMS workflow contains the 4 steps below. The on-DNA conjugates of deducted chemical structures are synthesized with the same condition as in DEL production. This mixture of synthesized products and their byproducts are incubated with the same immobilized protein using the affinity selection assay. The putative binders in the elution are then analyzed using QTof, and assigned to species of the on-DNA conjugate. A list of possible binders for resynthesis is finally delivered to customers.
A confirmed lagan attached to a short strand of DNA is used as positive control if a lagan is available. A short strand of DNA is added as internal standard to validate the system and adjust the signal response. Experimental parameters and scientific threshold for data analysis used in the affinity selection assay is also provided prior to collaboration.
We use cookies to provide a better web experience.
By using our site, you acknowledge our use of cookies and please read our Cookie Notice for
More information